Introduction¶
What is lib5c? How is it laid out? What can I do with it?
These are the questions that will be answered in this section.
What is lib5c?¶
lib5c is a comprehensive, modular library for analyzing the results of
Chromosome Conformation Capture Carbon Copy (5C) experiments.
How is lib5c organized?¶
lib5c is organized into three distinct layers:
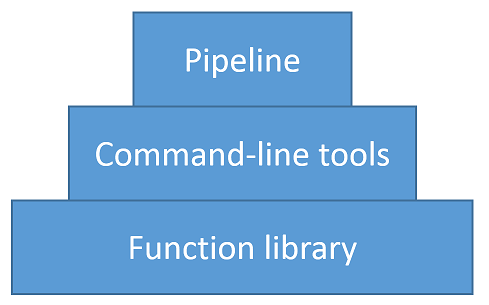
Each layer of the pyramid rests on the ones below it. At its base, lib5c is
a library of modular, reusable functions that can be used for various tasks in
5C analysis. A set of command-line tools rests on top of these functions,
making them easier to use for the most common use cases. The command-line tools
in turn support a pipeline, which provides a customizable interface for
performing entire sequences of analysis steps quickly and reproducibly.
Each of the three tutorials in the following sections of this documentation explain one of the three levels by example, starting from the top of the pyramid. You can start from the top, and go as deep as you feel comfortable going.
What can I do with lib5c?¶
This list shows just a few of the many things you can do with lib5c.
Analyze:
Correct for locus-specific bias factors.
Bin or smooth fragment-level interaction data.
Construct distance-dependence models.
Perform statistical modeling.
Classify significant interactions from a two-condition experiment.
Visualize:
Draw contact frequency heatmaps to see the architecture of the genome.
Draw bias factor heatmaps to idenfity covariates in the interaction data.
Visualize expected models and variance estimates.
Compare theoretical distributions to the real data.
Visualize enrichments between interaction classes and traditional genome annotations.